Introduction
Cancer cells rewire their metabolism and the glycolytic metabolic phenotype was the first described [1]. Depending on the cellular context, cancer cells can manifest different metabolic phenotypes, ranging from the glycolytic phenotype to a glycolytic partial phosphorylative and/or phosphorylative phenotype [2]. Furthermore, metabolic status in cancer cells may depend on the relative contributions of different oncogenic and tumor-suppressor genes [3].
Sex steroid hormones, including Estradiol (E2) and progesterone (OHPg), exerta wide variety of physiological and pathological actions, by binding to their specific receptors [4-6]. For instance, the 21-carbon steroid OHPg binding to Progesterone Receptors (PRs) is able to induce cellular processes required for the development of normal mammary gland, but also in breast cancer [7]. In mouse model, OHPg stimulates normal human breast epithelium through a paracrine mechanism promoting pre-neoplastic progression [8]. Human PRs derive from a single gene as two isoforms, PR-A and PR-B and their ratio varies in different tissues and patho-physiological conditions, as in breast malignancy [9]. Particularly, the ratio between PR-A and PR-B is augmented in breast tumors from patients with poor prognosis, suggesting a protective role of PR-B in breast cancer [10-12]. The level of these steroid hormone receptors in patients with breast cancer has been used in clinical management as an indicator of endocrine responsiveness [13]. Approximately 75 % of primary breast cancers express ER, and more than half also express PR [14]. Although, the clinical significance of ER evaluation has been well established, the role of PR remains controversial, and whether PR assessment is necessary has been debated for years [15].
In our recent paper, we reported in breast cancer a functional crosstalk between PR-B and PTEN in inducing autophagy through the involvement of AKT activation [16]. Autophagy is retained a possible way to reprogram cellular metabolism by activated oncogenes and inactivated tumor suppressors.
The survival of cancer cells is mainly dependent on their metabolic reprogramming and alteration of tumour metabolism represents a truthful approach to induce cancer cell death. To identify a cancer metabolic phenotype, it is needed to understand how the metabolic pathways are arranged in comparison with normal metabolism. In this context, our is the first study evidencing the OHPg/PR-B involvement on tumor cell metabolism, at our knowledge.
In order to investigate other tumour-suppressive pathways through which OHPg/PR-B signalling carries out a protective action in breast cancer, we evaluated its possible action on the alteration of breast cancer cell lines metabolic reprogramming.
Materials and methods
Chemicals and assay kits: All chemicals, including 17-Hydroxyprogesterone (OHPg), were bought from Merk Life Sciences (Milan, Italy). Glucose-6-phosphate dehydrogenase (G6PDH), Lactate Dehydrogenase (LDH) and lipase activities, as well as glucose and triglycerides assays kits were from Inter-Medical (Biogemina Italia Srl, Catania, Italy). Molecular Probes ATP Determination Kit (A22066) was obtained from Thermo Fisher Scientific (Milan, Italy).
Cell cultures: Human breast cancer cells MCF7 and T47D were maintained according to supplier’s recommendations (ATCC). MCF7 and T47D were cultured respectively in DMEM/F-12 and RPMI 1640 media, containing 5% FBS, 1% L-glutamine, 1% Eagle’s nonessential amino acids and 1 mg/ml penicillin/streptomycin (Merk Life Sciences, Milan, Italy)in a 5% CO2 humidified atmosphere. Every 4 months, cells were authenticated by single tandem repeat analysis at our Sequencing Core and tested for morphology, doubling times, and mycoplasma negativity, using MycoAlert Mycoplasma Detection Assay (Lonza, Rome, Italy). Twenty-four hours (h) before each experiment, cells were shifted in serumfree medium and treated with different concentrations of OHPg in medium containing 5% charcoal-treated FCS to reduce the endogenous steroids concentration. OHPg concentrations were chosen on the basis of our previous studies [4]. Furthermore, since in the major part of the experiments the 10 nM OHPg was the more efficacious in inducing an effect (probably since this is within the physiological range in women), we referred particularly to this concentration, although in some experiments we also presented three different doses.
Seahorse XFe96 metabolic flux analysis: Oxygen Consumption Rates (OCRs) in real times and Extracellular Acidification Rates (ECAR), which allow to assess the cellular respiration, glycolysis and ATP production in live cells, were determined using the Seahorse Extracellular Flux (XFe96) analyzer (Seahorse Bioscience, USA). Briefly, 1 × 104 cells (MCF7 and T47D) per well were seeded into XFe96 well cell culture plates in presence of DMEM/F-12 and incubated for 24h to allow cell attachment. After which, cells were starved in serum free media for 24h and successively treated in medium containing 5% charcoal-treated FCS, with OHPg at different concentrations. After 48h of treatment, cells were washed in pre-warmed XF assay media, supplemented with 10 mM glucose, 1 mM Pyruvate, 2 mM L-glutamine, and adjusted at 7.4 pH (only for OCR measurement). Cells were then maintained in 175 μL/ well of XF assay media at 37°C, in a non-CO2 incubator for 1 h. In to the sensor cartridge of XFe96, during the incubation time, were loaded 25 μLof XF assay media containing 80 mM glucose, 9 μM oligomycin, and 1M 2-deoxyglucose, for ECAR measurement and 25 μL of XF assay media with 10 μM oligomycin, 9 μM FCCP, 10 μM rotenone, 10 μM antimycin A for OCR determination. Measures were normalized by protein content (SRB assay).
Western blotting: Cells were treated as indicated and total protein extracts were obtained as previously described [17]. RIPA buffer (50 mM Tris–HCl, pH 7.4, 150 mM NaCl, 1% NP-40, 0.25% Na deoxycholate and specific inhibitors) was used for cell lysis and protein content was determined by Bradford dye reagent (BioRad). Equal amounts of protein extracts were resolved on a 10% Sodium Dodecyl Sulfate-Polyacrylamide Gel (SDS-PAGE), transferred to nitrocellulose membranes (Amersham Biosciences, Italy), probed overnight at 4°C with antibodies against: PFK1 (G-11 sc166722), PDH-E1α(D-6 sc-377092), αKGD (C-20 sc-49589), ACLY (5F8D11 sc-517267), FAS (H-300 sc 20140), PR (F-4sc166169) (Santa Cruz Biotechnology DBA, Italy), ACC (#3662)PDK (#3062) (Cell Signalling Technologies, Euroclone, Italy). β-actin (C-4, sc-47778) (Santa Cruz Biotechnology DBA, Italy) was used as loading control. Proteins were detected by horseradish peroxidase-linked secondary antibodies (DBA, Milan, Italy) and revealed using the ECL Plus Western Blotting detection system (Amersham Pharmacia Biotech, United Kingdom). Odyssey FC (Licor, Lincoln, NE, USA) and Scion Image laser densitometry scanning program were used, respectively, for acquisition and quantification of each band of interest.
Lipid-mediated transfection of siRNA duplexes: Eighty percent confluent cells was transfected with functionally verified siRNA directed against human PR-B or with a control siRNA (Qiagen, MI, Italy) that did not match with any human mRNA used as a control for non-sequence specific effects (NS siRNA). Cells were transfected using Lipofectamine 2000 reagent (Invitrogen, Paisley, UK) according to the manufacturer’s instructions as previously reported [17] and then treated as indicated.
Lactate dehydrogenase activity: LDH activity, showed as absorbance change at 340 nm, was evaluated on cell lysates as previously indicated [18].
Glucose 6 phosphate dehydrogenase activity: G6PDH enzymatic activity was measured by analysing the increase of absorbance at 340 nm [18]. Data have been presented as nM/min/µg protein.
ATP assay: Quantitative determination of ATP was measured by a bioluminescence assay with recombinant firefly luciferase and its substrate D-luciferin (light emission at 560 nm at pH 7.8) as previously described [18]. Data are presented as nM/µg protein.
Isocitrate dehydrogenase activity assay: Isocitrate Dehydrogenase (ICDH) activity was measured as previously reported [18]. The reaction was performed on the cell lysate using a final 1 ml reaction volume containing 0.2 mg protein, 50 mM Tris-Cl pH 7.4, 5 mM MnCl2, 0.25 mM NADP+, 0.25 mM isocitrate, at 37°C. Reaction was monitored at 340 nm for 2 minutes (min); NADPH production was calculated using an NADPH extinction coefficient of 6.26103 M-1 cm-1. Data are presented as nM/min/µg protein.
Malic enzyme activity assay: Maleic Acid (ME) activity was determined as previously reported [18], using a final 1 ml reaction volume containing 0.3 mg protein, 50 mM Tris-HCl pH 7.4, 5 mM MnCl2, 0.1 mM NADP+, 5 mM malate, at room temperature. Reaction was checked at 340 nm for 2 min; the NADPH production was calculated using an NADPH extinction coefficient of 6.26103 M-1 cm-1. Data are presented as nM/min/µg protein.
Triglycerides assay: Triglycerides content were evaluated in duplicate by a GPO-POD enzymatic colorimetric method according to manufacturer’s instructions as previously described [18]. Data are presented as nM/µg protein.
Lipase activity assay: Lipase activity was measured, by Panteghini et al. [19] method, based on the use of 1,2-o-dilauryl-rac-glycero-3-glutaric acid-(6’-methylresorufin) ester as substrate, as previously described [18]. Data are presented as nM/min/µg protein.
Octanoyl CoA dehydrogenase activity assay: Octanoyl CoA dehydrogenases catalyze the initial step in each cycle of fatty acid β-oxidation. 50 µg of protein lysates was added to a buffer containing 20 mm Mops, 0.5 mm EDTA and 100 lm FAD at pH 7.2. Reduction of FAD to FADH2 was checked at 340 nm after addition of octanoyl-CoA (100 µg M) every 20s for 1.5 minutes. Data are expressed as nmol⁄min⁄µg protein. The enzymatic activity was determined with three control media: one without octanoyl-CoA as substrate, another without the coenzyme (FAD) and the third without either substrate or coenzyme (data not shown).
Statistical analysis: Data obtained by Seahorse XFe96 were analysed by XFe96 and GraphPad Prism softwares, using one-way ANOVA and Student’s t-test. All experiments were performed in quintuplicate, three times independently. Data obtained from the enzymatic activities, as well as, glucose and triglycerides quantification assays (six independent experiments using duplicate determinations), were presented as the mean ± SEM. The Western blot analyses were performed in at least four independent experiments. Western blotting densitometric analyses, indicated as the numbers above blot bands, represents the mean intensities of the bands evaluated in terms of arbitrary densitometric units. The differences in mean values were calculated using Analysis of Variance. The Wilcoxon test was used after analysis of variance as post hoc test. A value of P ≤ 0.05 was retained statistically significant.
Results
OHPg modulate energy metabolismin MCF7 and T47D cells
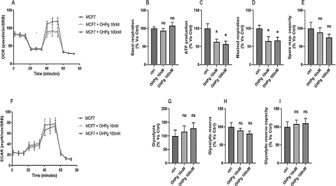
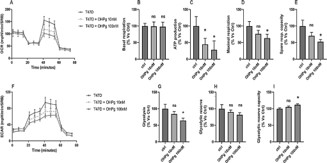
Under typical in vitro cell culture conditions, OCR is an indicator of mitochondrial respiration, while ECAR is predominantly a measure of lactic acid formed during glycolytic way. Measuring OCR and ECAR simultaneously, may be indicative of the dynamic interplay between glycolysis and oxidative metabolism [20]. To see metabolic adaptation dynamically and on live cells, we have evaluated the metabolic profile in MCF7 and T47D cells treated or not for 48h with OHPg 10 nM and 100 nM (Figure 1 and Figure 2). In MCF7 cells both OHPg concentrations reduced the OCR amount, as well as the ATP production and maximal respiration levels (Figure 1A, C and D), but increased ECAR values (Figure 1F). In the contrary, not significant changes were founded in basal respiration, spare respiratory capacity, glycolysis, glycolytic reserve and glycolytic reserve capacity levels (Figure 1B, E, G,H and I). Regarding T47D cells, OHPg 10 nM and 100 nM decreased OCR levels, ATP production and ECAR namounts (Figure 2A, C and F). Of note, only the OHPg 100 nM treatment reduced the maximal respiration, spare respiratory capacity and glycolysis (Figure 2D, E and G) and increased glycolytic reserve capacity (Figure 2I).
OHPg/PR-B regulated glycolysis and intracellular LDH activity in breast cancer cells
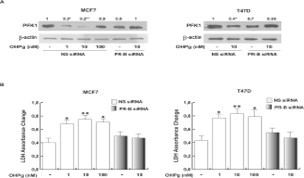
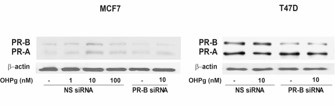
To better define how OHPg/PR-B could regulate the glycolytic pathways in cancer cells, we determined the Phosphofructokinase-1 (PFK1) expression and intracellular LDH activity in MCF7 and T47D cells treated with increasing OHPg concentrations for 24 h. In both cell lines, OHPg reduced PFK1 expression (Figure 3A). LDH enzymatic activity upon OHPg stimulation was augmented (Figure 3B). These effects were abrogated transfecting in MCF7 and T47D cancer cells a specific PR-B siRNA (Figure 3 A,B and Supplementary Figure 1), hence suggesting the PR-B involvement in these actions induced by OHPg.
OHP/PR-B effects on biosynthetic contributions in breast cancer cells
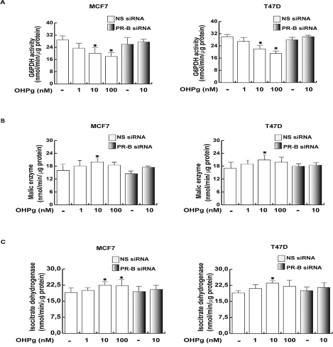
Glucose is also engaged through the Pentose Phosphate Pathway (PPP) which is up regulated in many types of tumors and the first enzyme of the pathway is the G6PDH [21]. In both cell lines, OHPg treatment determined a significant decline of the G6PDH activity at OHPg 10 and 100 nM (Figure 4A), addressing how the lowering of G6PDH in treated-cells also contributed to the decrease of glucose utilization. To better define the OHPg/ PR-B action on biosynthetic state of our cells, we investigated the activity of other two enzymes involved in the production of NADPH, such as ME and ICDH. In MCF7 and T47D cells, both enzymatic activities were increased by OHPg 10 nM through the involvement of PR-B (Figure 4B,C).
OHPg/PR-B modulate bioenergetic requirements in breast cancer cells, inducing a ‘truncated’ TCA and altering the de novo fatty acid synthesis.
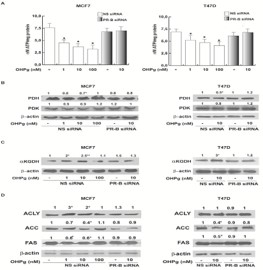
Cancer cells constantly require high amount of ATP [22]. To confirm our results obtained by Seahorse analysis and to evaluate the role of PR-B/OHPg in ATP content, we performed a bioluminescence assay to quantify the ATP levels in our cell lines upon OHPg treatments in combination or not with PR-B siRNA. All the OHPg concentrations used caused a reduction of ATP content and these effects were abrogated by transfecting PR-B siRNA (Figure 5A). Pyruvate is transported to the mitochondria to be oxidized by the Pyruvate Dehydrogenase (PDH), which in turn is regulated by the pyruvate dehydrogenase kinase (PDK) [23]. As shown in Figure 5A, in both cell lines the PDH expression was significantly reduced after OHPg 10 nM treatment and this effect was abrogated by silencing PR-B. OHPg treatment did not influenced PDK content in both cell lines (Figure 5B). Moreover, OHPg 10nM inducedthe α-ketoglutarate dehydrogenase (αKGDH) expression through the involvement of PR-B (Figure 5C). These results suggest that OHPg may contribute to refuel the TCA cycle at level of the αKGDH, hence inducing in breast cancer cells a ‘truncated’ TCA cycle, which can include spill of citrate to the cytosol for lipid synthesis [24]. Interestingly, we observed that only in MCF7 cancer cells the ATP Citrate Lyase (ACLY) content increased after OHPg 1 and 10 nM treatment via PR-B (Figure 5D), while in T47D, OHPg stimulation did not have any effect. Conversely, in both cell lines OHPg decreased the Acetyl-CoA Carboxylase (ACC) and Fat Acid Synthase (FAS) expression which were rescued transfecting PR-B-siRNA (Figure 5D).
OHPg/PR-B promoted a lipid lowering effect in breast cancer cells
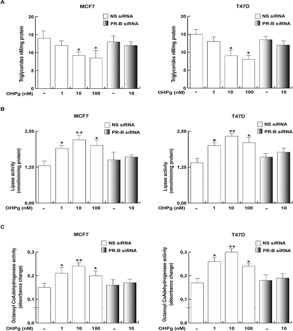
Similar to glucose, fatty acid metabolism supports both the biosynthetic and the bioenergetics needs for cell growth and survival [25]. OHPg-stimulated MCF7 as well as T47D cells through PR-B diminished the triglycerides content versus untreated cells (Figure 6A). Concomitantly, the lipase and Octanoyl CoA-dehydrogenase activities, both fatty acid catabolic processes, increased at all the OHPg concentrations used (Figure 6B, C respectively). The PR-B-siRNA counteracted the OHPg effects (Figure 6B, C).
Discussion
Cancers are extremely heterogeneous diseases with individual metabolic features. Nevertheless, aerobic glycolysis is often found in malignant tumors, OXPHOS still contributes to energy production [26]. Therefore, a multifaceted approach is important to define the metabolic phenotype of a cancer. Herein, we evidenced various features of tumor metabolism influenced by OHPg/PR-B in human breast cancer cells, unveiling a novel protective action for OHPg/PR-B. Both in MCF7 and T47D cells, first we studied the metabolic phenotype by the Seahorse XFe96 Analyser. From the data obtained, the OHPg did not influence glycolysis way, although in T47D a reduction at OHPg 100 nM was observed. The ATP levels were significantly reduced after OHPg treatment in both cell lines. To better define the metabolism of our cell lines, we analyzed different key factors, which characterizes both glucose metabolism and ATP production. By evaluating glucose metabolism in MCF7 and T47D cells upon increasing OHPg treatments, we observed that the PFK1 expression was eminently decreased, concomitantly the intracellular LDH activity increased. LDH, a long-lived protein is degraded mainly or exclusively by autophagy. In this context, the present results well fit with our previous findings indicating that OHPg through PR-B/Bcl-2 axis drives an impaired autophagy that facilitates the irreversible growth arrest determining breast cancer cell fate [11,16]. Lactate synthesis can exert a metabolic control over glycolysis, through the inhibition of PFK1 enzyme, a rate limiting step of the glycolytic pathway [27]. Recently, Mulukutla and co-authors showed lactate inhibitory effect on PFK1, which in turn results in slowing down the glycolytic flux [28]. It is known that the inhibition of PFK1 determines the accumulation of fructose-6-phosphate, and then isomerized to glucose-6-phosphate fueling also the PPP. This pathway is crucial to sustain antioxidant and anabolic functions in many cancer cells [29]. Our results showed that OHPg/PR-B reduce the G6PDH activity thus disturbing the PPP rate. Altogether, these data are in agreement with another our recent study, where we demonstrated that OHPg/PR-B induced the autophagy process through PTEN, causing an inhibition of the AKT signal [16] which is a glycolysis stimulator, therefore the down regulation of its signaling cascade can also explain the low glycolytic flux as it emerged from this finding.
Other than the PPP, the cytosolic enzymes ME and ICDH contribute to the production of NADPH [30], interestingly both enzymatic activities increased after OHPg stimulation in our cell models. Pyruvate may be reduced to lactate and moved out of the cell or to go into the mitochondria to be oxidized by the PDH into AcetylCoA [23].
PDH and KGDH complexes regulate most of the carbon flow into the TCA cycle, derived from pyruvate and other sources respectively [31]. Our data showed that the refueling of TCA cycle may occur prevalently through the KGDH, since its expression increased, while that of PDH was reduced by OHPg. The heightened need in cancer cells for biosynthetic intermediates may also result in the augmented use of a ‘truncated’ TCA cycle. Unfortunately, a persisted use of a truncated TCA cycle alters the integrity of mitochondrial function, accordingly your data that showed a strong reduction of the ATP content, using both the Seahorse analyzer and a bioluminescence assay.
A shortened TCA cycle includes the citrate extrusion in the cytosol to be used for fatty acid de novo synthesis [32]. In fact, citrate may be cut by ACLY into OAA and Acetyl-CoA. The latter may be used for de novo lipid synthesis, a complex process involving enzymes such as ACC and FAS. In our study, OHPg increased ACLY expression, whereas ACC and FAS levels were reduced, and this may imply a switch towards the OAA cytosolic formation, which in turn through ME and ICDH activities induced by OHPg, sustained pyruvate and thus LDH activity. These data are also in agreement with the increased intracellular LDH activity as we found. Furthermore, OHPg induced a general lipid lowering effect since de novo fatty acid synthesis diminished. In fact, triglyceride levels decreased, while concomitantly the lipase and the FAO activities increased. Fatty acids are an important energy source that can produce more ATP than carbohydrates when required. These observations and our data suggest that OHPg/PR-B may induce the lipid degradation in breast cancer cells to recover energy, failing however in their intent.
Collectively, our findings indicate for the first time that activation of the PR-B by its own natural ligand influences several metabolic pathways in breast cancer cells, reducing both glucose, oxidative and lipid metabolism. Herein, we discovered another distinctive protective action of OHPg/PRB against breast cancer. We hypothesize that OHPg may be taken into account, in combination with other types of targeted chemotherapy, to improve breast cancer treatment outcomes.
Conclusion
PR-B is a key prognostic marker for breast cancer patients, up to date it is underexplored. OHPg through PR-B disrupts metabolic reprogramming in breast cancer cells, altering glycolysis, oxidative phosphorylation and lipogenesis, hence OHPg can be considered in chemotherapy to improve cancer treatment outcomes.
Declarations
Conflicts of interests: The authors declare no conflict of interest.
Funding sources: This work was supported by MinisteroIstruzione Università e Ricerca (ex 60 % 2020).
Acknowledgments: We would like also to thank Serena and Maria Clelia Gervasi for the English language review of the manuscript.
References
- Sun L, Suo C, Li ST, Zhang H, Gao P. Metabolic reprogramming for cancer cells and their microenvironment: Beyond the Warburg Effect. Biochim Biophys Acta Rev Cancer. 2018; 1870; 51-66.
- Pike WL, Wu M. Rapid Analysis of Glycolytic and Oxidative Substrate Flux of Cancer Cells in a Microplate. PLoS One. 2014; 10: e109916.
- Pavlova NN, Zhu J, Thompson CB. The hallmarks of cancer metabolism: Still emerging. Cell Metab. 2022; 34: 355-377.
- De Amicis F, Russo A, Avena P, Santoro M, Vivacqua A, et al. In vitro mechanism of action for down-regulation of ERalpha expression by epigallocatechin gallate in ER+/PR+ human breast cancer cells. Mol Nutr Food Res. 2013; 57: 840-853.
- De Amicis F, Guido C, Perrotta I, Avena P, Panza S, et al. Conventional progesterone receptors (PR) B and PRA are expressed in human spermatozoa and may be involved in the pathophysiology of varicocoele: a role for progesterone in metabolism. Int J Androl. 2011; 34: 430-445.
- Hilton HN, Clarke CL, Graham JD. Estrogen and progesterone signalling in the normal breast and its implications for cancer development. Mol Cell Endocrinol. 2018; 5: 2-14.
- Horwitz KB, Sartorius CA. 90 YEARS OF PROGESTERONE. Progesterone and progesterone receptors in breast cancer: past, present, future. J Mol Endocrinol. 2020; 65: T49-T63.
- Trabert B, Sherman ME, Kannan N, Stanczyk FZ. Progesterone and Breast Cancer. Endocr Rev. 2020; 4: 320-344.
- Allison KH, Hammond MEH, Dowsett M, McKernin SE, Carey LA, et al. Estrogen and Progesterone Receptor Testing in Breast Cancer: ASCO/CAP Guideline Update. J Clin Oncol 2020; 38: 1346-1366.
- De Amicis F, Zupo S, Panno ML, Malivindi R, Giordano F, et al. Progesterone Receptor B recruits a repressor complex to a half-PRE site of the Estrogen Receptor alfa gene promoter. Mol Endocrinol. 2009; 23: 454-465.
- De Amicis F, Guido C, Santoro M, Giordano F, Donà A, et al. Ligand activated progesterone receptor B drives autophagy-senescence transition through a Beclin-1/Bcl-2 dependent mechanism in human breast cancer cells. Oncotarget. 2016; 7: 57955-57969.
- Montalto FI, Giordano F, Chiodo C, Marsico S, Mauro L, et al. Progesterone Receptor B signaling Reduces Breast Cancer Cell Aggressiveness: Role of Cyclin-D1/Cdk4 Mediating Paxillin Phosphorylation. Cancers (Basel). 2019; 11: 1201.
- Giulianelli S, Lamb CA, Lanari C. Progesterone receptors in normal breast development and breast cancer. Essays Biochem. 2021; 65: 951-969.
- Dai X, Xiang L, Li T, Bai ZJ. Cancer Hallmarks, Biomarkers and Breast Cancer Molecular Subtypes. Cancer. 2016; 7: 1281-1294.
- Onitilo AA, Engel J, Joseph AO, Li YH. Is oestrogen receptor-negative/progesterone receptor-positive (ER-/PR+) a real pathological entity? Ecancermedicalscience. 2021; 15: 1278.
- De Amicis F, Guido C, Santoro M, LanzinoM, Panza S, et al. A novel functional interplay between Progesterone Receptor-B and PTEN, via AKT, modulates autophagy in breast cancer cells. J Cell Mol Med. 2014; 18: 2252-2265.
- Panza S, Santoro M, De Amicis F, Morelli C, Passarelli V, et al. Estradiol via estrogen receptor beta influences ROS levels through the transcriptional regulation of SIRT3 in human seminoma TCam-2 cells. Tumour Biol. 2017; 39: 101.
- Santoro M, Guido C, De Amicis F, Sisci D, Cione E, et al. Bergapten induces metabolic reprogramming in breast cancer cells. Oncology Reports. 2016; 35: 568-576.
- PanteghiniM, Bonora R, Pagani F. Measurement of pancreatic lipase activity in serum by a kinetic colorimetric assay using a new chromogenic substrate. Ann Clin Biochem. 2001; 38: 365-370.
- Zhang J, Zhang Q. Using Seahorse Machine to Measure OCR and ECAR in Cancer Cells. Methods Mol Biol. 2019; 1928: 353-363.
- Cho ES, Cha YH, Kim HS, Kim NH, Yook JI. The Pentose Phosphate Pathway as a Potential Target for Cancer Therapy. Biomol Ther (Seoul). 2018; 26: 29-38.
- Phan LM, Yeung SC, Lee MH. Cancer metabolic reprogramming: importance, main features, and potentials for precise targeted anti-cancer therapies. Cancer Biol Med. 2014; 11: 1-19.
- McCommis KS, Finck BN. Mitochondrial pyruvate transport: a historical perspective and future research directions. Biochem J. 2015; 466: 443-454.
- Icard P, Coquerel A, Wu Z, Gligorov J, Fuks D, et al. Understanding the Central Role of Citrate in the Metabolism of Cancer Cells and Tumors: An Update. Int J Mol Sci. 2021; 22: 6587.
- Zhu J, Thompson CB. Metabolic regulation of cell growth and proliferation. Nat Rev Mol Cell Biol. 2019; 20: 436-450.
- Slavov N, Budnik B, Schwab D, Airoldi E, Van Oudenaarden A. Constant growth rate can be supported by decreasing energy flux and increasing aerobic glycolysis. Cell Rep. 2014; 7: 705-714.
- Leite TC, Coelho RG, Da Silva D, Coelho WS, Marinho-Carvalho MM, et al. Lactate downregulates the glycolytic enzymes hexokinase and phosphofructokinase in diverse tissues from mice. FEBS Lett. 2011; 585: 92-98.
- Mulukutla BC, Yongky A, Daoutidis P, Hu WS. Bistability in glycolysis pathway as a physiological switch in energy metabolism. PLoS One. 2014; 9: e98756.
- De Berardinis RJ, Chandel Navdeep S. Fundamentals of cancer metabolism. Sci Adv. 2016; 2: e1600200.
- Ren JG, Seth P, Clish CB, Lorkiewicz PK, Higashi RM, et al. Knockdown of malic enzyme 2 suppresses lung tumor growth, induces differentiation and impacts PI3K/AKT signaling. Sci Rep 2014; 4: 5414.
- Mullen AR, Hu Z, Shi X, Jiang L, Boroughs LK, et al. Oxidation of alpha-ketoglutarate is required for reductive carboxylation in cancer cells with mitochondrial defects. Cell Rep. 2014; 7: 1679-1690.
- Lussey-Lepoutre C, Hollinshead KE, Ludwig C, Menara M, Morin A, et al. Loss of succinate dehydrogenase activity results in dependency on pyruvate carboxylation for cellular anabolism. Nat Commun. 2015; 6: 8784.